Joint efforts combine a total of 24 petaflops of GPU computing power between ImmunityBio and Microsoft, enabling simulation of 200 μs per day of binding between the COVID-19 spike protein and the host ACE-2 surface protein – a crucial step to develop proteins to stop the infection
EL SEGUNDO, Calif., April 1, 2020 – ImmunityBio, Inc., a privately held immunotherapy company within the NantWorks ecosystem of companies, today announced it is collaborating with Microsoft to leverage the company’s Azure platform to perform a highly detailed computational analysis of the spike protein structure of the SARS-CoV-2, the novel coronavirus responsible for the global pandemic. The spike protein serves as the “doorway” for the virus to enter human cells, making it a high-priority target for a vaccine or antibody therapies to fight the virus.
A digital blueprint of the spike protein obtained via a process called cryo-electron microscopy was published in February by researchers at the University of Texas and the National Institutes of Health. The ImmunityBio and Microsoft teams have taken that one step further by applying a technique called molecular dynamics to the blueprint. Molecular dynamics analyzes the physical movements of the virus components at the atomic level over an extended period of time and runs a series of computationally intensive simulations that result in a detailed model of the most likely solution structure of the spike protein.
Having a detailed model of the spike protein complex is crucial for researchers seeking to develop effective vaccines or therapies. The protein is the key to the mechanism the virus uses to invade cells in the body and cause an infection. The spike protein, so called because it protrudes from the surface of the viral particle, binds to the ACE2 receptor on the surface of an epithelial cell in the human respiratory tract. Once it has done so, the genetic material from the virus is able to enter the cell and commandeer its function so the cell produces copies of the virus in large numbers.
The human immune system normally attempts to fight these infections by creating antibodies that recognize the protein, specifically neutralize it and thus keep the cellular “doorway” closed. Because the SARS-CoV-2 virus is novel – that is, not seen previously in humans – the immune system in most people is unable to develop antibody resistance rapidly enough to keep ahead of the infection.
Both ImmunityBio and Microsoft donated their massive networked computing power and advanced algorithms needed to derive the model in days, rather than the months it would normally require using older technical approaches. With this model in hand, researchers working on vaccines and treatments have a clear therapeutic target that will streamline their work in finding ways to treat the pandemic.
“The preclinical process of finding and selecting a target for a traditional therapy can take years, which we don’t have in our fight against the coronavirus,” said Dr. Patrick Soon-Shiong, Chairman and CEO of ImmunityBio, Inc. “Across our portfolio of biotech companies, including ImmunityBio and NantKwest, we are committed to helping find effective therapeutics for coronavirus and other infectious diseases. Association of the COVID-19 spike protein with host ACE-2 surface proteins is a crucial step in infection. Structures of this complex are available, but understanding how the two proteins dynamically interact is critical to targeting it. This gives us valuable information about how COVID-19 binds to lung cells and what drives the association. The involvement of Microsoft and its abundant computing infrastructure will bolster our drug discovery and development progress by our computer scientists and molecular modelers towards entering an optimal therapeutic candidate in clinical trials this year.”
“Microsoft is committed to bring our technology and expertise to bear in solving the complex computing problem of modeling this protein,” said Peter Lee, Corporate Vice President, Microsoft AI & Research. “With ImmunityBio we are working to speed the effort to find a treatment for this deadly virus that has affected every part of the globe.”
Microsoft, collaborating with ImmunityBio’s engineers and scientists quickly deployed a High Performance Compute cluster on Microsoft Azure cloud services. The cluster contains over 1,250 NVIDIA V100 Tensor Core high performance graphics processing units (GPUs) specifically designed for machine learning and other computationally intensive applications. Similarly, ImmunityBio has deployed its 320 GPU cluster, which has always been optimized for and dedicated to molecular modeling of proteins, antibodies, antivirals, and targeted small molecule drugs.
“Our joint efforts between Microsoft and ImmunityBio bring together an incredible amount of computing power to help create models for researchers working on vaccines and therapeutics,” said Dr. James Weinstein, Senior Vice President, Microsoft Healthcare. “We are pleased to support ImmunityBio and NantWorks to jointly find a path to end this pandemic.”
In 2011, Dr. Soon-Shiong and his team assumed control of the National LambdaRail, a 12,000-mile-long, high-speed national computer network that was used by the U.S. research and education communities and NASA to establish a federated super-computing cloud. Soon-Shiong has since expanded this cloud infrastructure to undertake molecular modeling of protein-to-protein docking and high affinity binding dynamics and he and the NANT team have successfully identified unique binding sites in important cancer-related proteins such as KRAS and Neoepitopes.
“With the advent of this pandemic, we have allocated our computing resources and our scientific skill sets to model the dynamics of the spike protein and its interaction with ACE 2. We are extremely grateful to Dr. Lee, Dr. Weinstein and their teams at Microsoft in supporting our efforts to discover novel binding sites to fight this war,” added Dr. Soon- Shiong.
About ImmunityBio
ImmunityBio, Inc. is a privately held immunotherapy company with a broad portfolio of biological molecules at clinical stages of development. The Company’s goals are to employ this portfolio to activate endogenous Natural Killer (NK) and CD8+ T cells in the fields of cancer and infectious disease. Specifically, in regards to cancer, ImmunityBio’s goal is to develop a memory T-cell cancer vaccine to combat multiple tumor types—without the use of high-dose chemotherapy. Regarding infectious disease, the Company is addressing HIV, influenza, and the coronavirus.
The Company’s first-in-human platform of technologies has enabled it to achieve one of the most comprehensive, late-stage clinical pipelines, activating both the innate (natural killer cell) and the adaptive immune systems. The product pipeline includes an albumin-linked chemotherapeutic (Aldoxorubicin), a novel IL-15 cytokine superagonist (N-803), checkpoint inhibitors, macrophage polarizing peptides, bi-specific fusion proteins targeting TGFb and IL-12, adenovirus, and yeast vaccine therapies targeting tumor associated antigens and neoepitopes.
In December 2019, the U.S. Food and Drug Administration (FDA) granted Breakthrough Therapy Designation to N-803 for BCG-unresponsive CIS non-muscle invasive bladder cancer (NMIBC). Other indications currently at registration-stage trials include BCG-unresponsive papillary bladder cancer, first and second-line lung cancer, and metastatic pancreatic cancer.
In the field of infectious disease, ImmunityBio’s goal is to develop therapies, including vaccines, for the prevention and treatment of HIV, influenza, and the coronavirus SARS-CoV-2.
Forward-Looking Statements
This press release contains forward-looking statements within the meaning of the Private Securities Litigation Reform Act of 1995. Forward-looking statements include statements concerning or implying that ImmunityBio will be successful in improving the treatment of the novel coronavirus. Risks and uncertainties related to this endeavor include, but are not limited to, the company’s beliefs regarding the success, cost and timing of its development activities and clinical trials.
Forward-looking statements are based on management’s current expectations and are subject to various risks and uncertainties that could cause actual results to differ materially and adversely from those expressed or implied by such forward-looking statements. Accordingly, these forward-looking statements do not constitute guarantees of future performance, and you are cautioned not to place undue reliance on these forward-looking statements. These forward-looking statements speak only as of the date hereof, and we disclaim any obligation to update these statements except as may be required by law.
Media Contact:
Jen Hodson
NANT
Jen@nant.com
562-397-3639
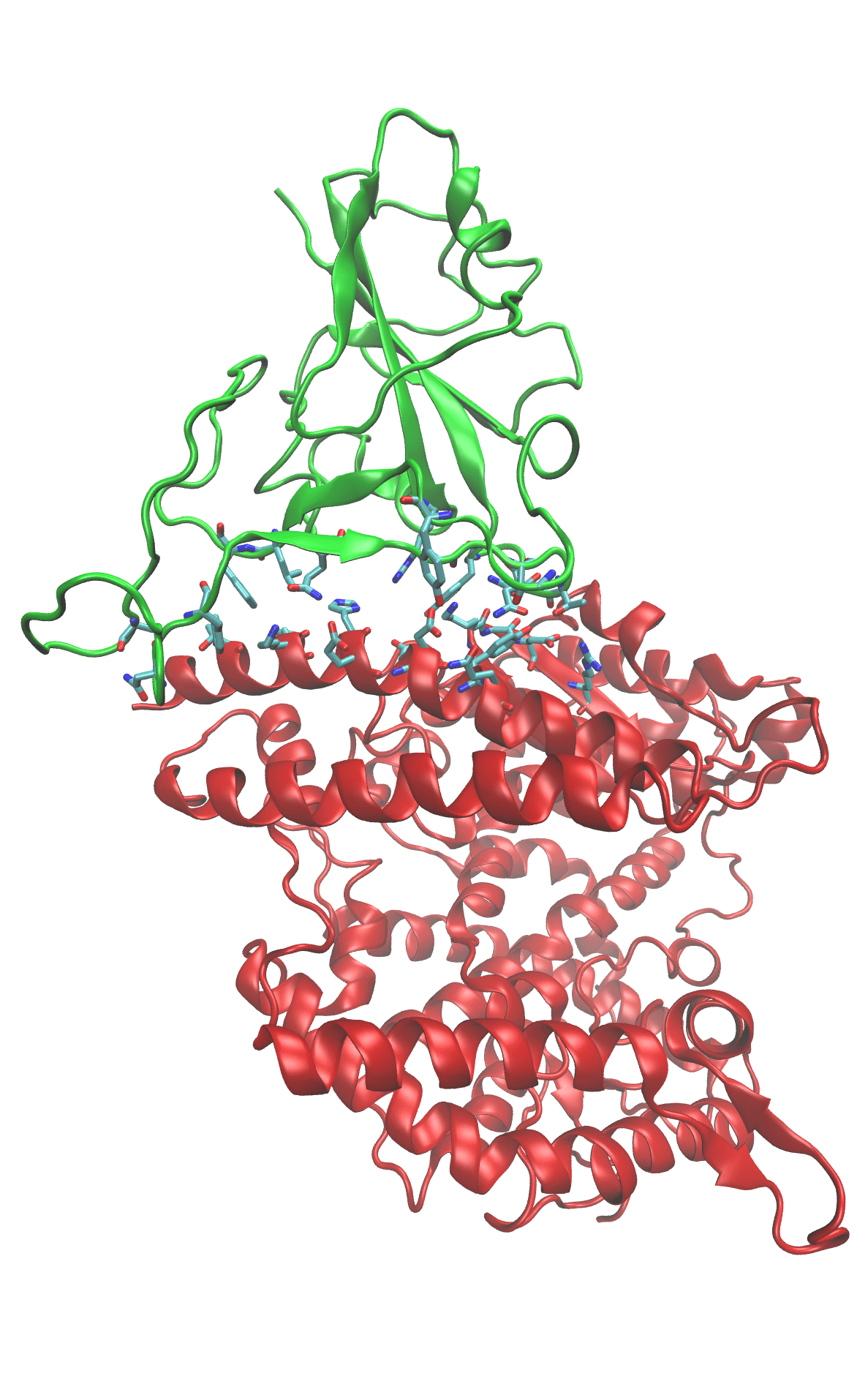
Binding between the SARS-COV-2 spike protein’s receptor binding domain (green ribbons) and the human ACE-2 receptor (red ribbons) is the first step in viral infection. The two proteins associate because their binding surfaces have complimentary shapes and charge distributions. The amino acids that make up the binding surfaces are shown as sticks and colored according to atom type. Dynamic simulations of the spike/ACE-2 complex let us monitor the individual interactions between amino acid residues. This atomic-scale description of binding shows which residues provide the driving force for binding. If that driving force can be interrupted, binding will be prevented and the infection stops.


